All Fragments Generated by Staggered Cut by Restriction Endonucleases Have
While there are hundreds of different restriction enzymes they all work in essentially the same way. Suppose that the disk of radius R shown is in equilibrium.

Type Iis Restriction Endonucleases Enable Gga A Type Ii Enzyme Ecori Download Scientific Diagram
Like all Type II restriction endonucleases.

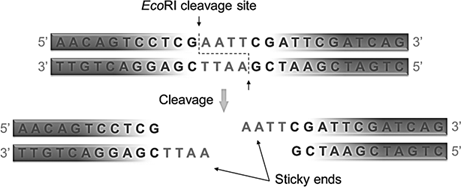
. Each DNA strand has overhanging and recessed ends. The cutting sites are frequently palindromic sequences sites at which the base sequence reading 5 to 3 on one strand is the same as the sequence reading 3 to 5 on the complementary strand of a. Why are blunt ends less useful.
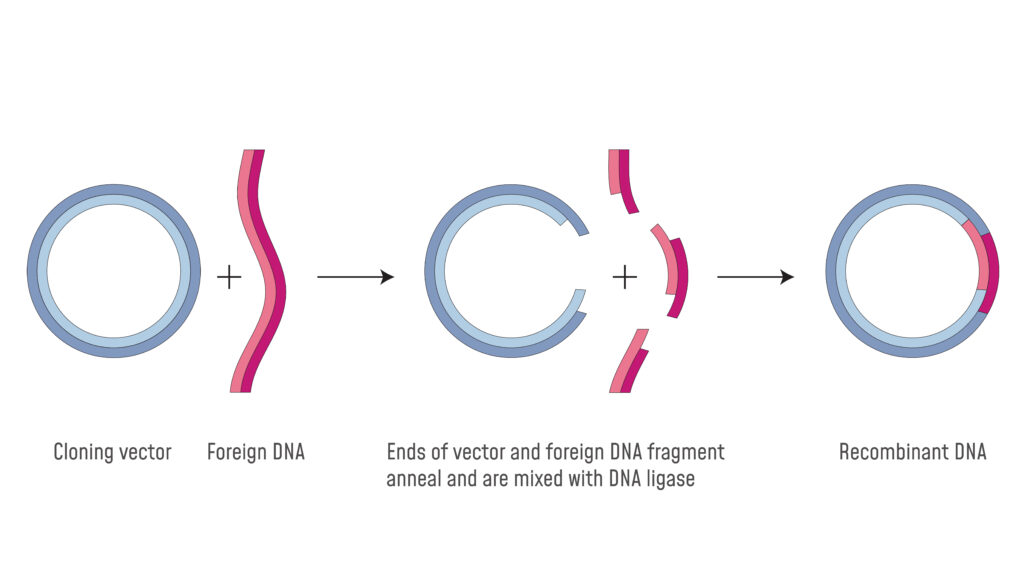
If two DNA molecules have matching ends they can be joined by the enzyme DNA ligase. All fragments generated by staggered cut by restriction endonucleases have complementary double-stranded ends. Sticky ends are generally more desired in cloning technology where a DNA ligase is used to join two DNA fragments into one because the yield.
Restriction enzymes are found in bacteria and other prokaryotes. Note that the incline has a coefficient of static friction μ s -μ Find the tension in the cord. A restriction enzyme that cuts the backbones of both strands at non-adjacent locations leaves a staggered cut generating two overlapping sticky ends.
All fragments cut by most restriction endonucleases have A complementary double-stranded ends. Staggered cuts are those in which the sites of cleavage on the two complementary DNA strands are not directly opposite each other as shown by the triangles in the example of Hindill above. Note that a consequence of staggered cuts is that.
Make staggered cuts at specific sequences in DNA in both donor DNA and plasmid. In the other style of cleavage by the restriction endonucleases the two strands of DNA are cut at two different points. Answer a is incorrect because the cleaving of DNA involves the use of restriction endonucleases to cut the DNA of the vector and the source DNA.
Whatever the starting material the result is a mixture of DNA fragments of a wide variety of lengths. Cohesive ends have unpaired DNA nucleotides on either 5- or 3- strand which are known as overhangs. Each enzyme recognizes one or a few target sequences and cuts DNA at or near those sequences.
HaeIII cuts both strands of DNA in the same location yielding restriction fragments with blunt ends. If DNA is digested by a restriction enzyme the DNA will be reduced to fragments of varying sizes depending on how many cutting sites for that restriction enzyme are present in the DNA. PvuII Haelll Alul are the examples of restriction endonucleases producing blunt ends.
The sticky ends aka. The recognition sites of number of type II restriction enzymes often make a staggered cut to leave molecule to generate short single-stranded ends. Many restriction enzymes make staggered cuts producing ends with single-stranded DNA overhangs.
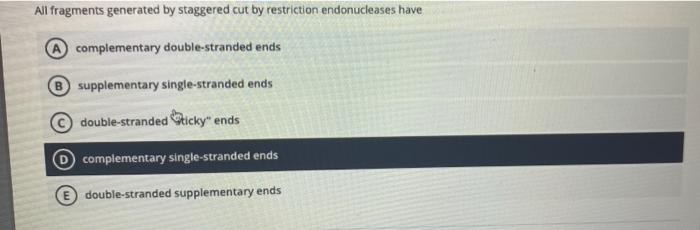
Many restriction enzymes make staggered cuts at or near their recognition sites producing ends with a single-stranded overhang. Many restriction endonucleases make staggered cuts in the two DNA strands such that the cut ends have a 2- or 4-base single-stranded overhang. All fragments generated by staggered cut by restriction endonucleases have complementary double-stranded ends B supplementary single-stranded ends double-stranded Sticky ends complementary single.
The remaining single stranded overhangs will be complimentary when cut by the same type of enzyme and can be easily ligated. DNA fragments generated by the restriction endonucleases in a chemical reaction can be separated by 1 centrifugation 2 polymerase chain reaction 3 electrophoresis 4 restriction mapping Practice questions MCQs Past Year Questions PYQs NCERT Questions Question Bank Class 11 and Class 12 Questions NCERT Exemplar Questions and PDF Questions with. These are called as sticky ends.
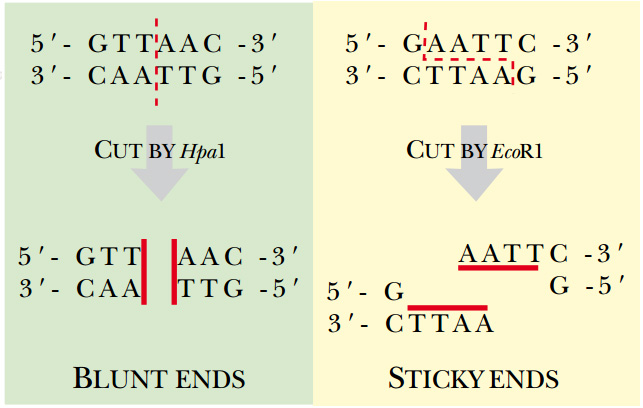
Some restriction enzymes cut in the middle of their recognition site creating blunt-ended DNA fragments. A straight cut of restriction enzymes generates blunt ends where both strands terminate in a base pair. Many restriction enzymes eg Eco RI Bam HI and Hind III make staggered single-stranded cuts producing short single-stranded projections at each end of the cleaved DNA called sticky ends.
The sticky ends aka. As a result the DNA ends generated by the enzymes shown here are staggered. These overhangs are most often generated by a staggered cut of restriction enzymes.
When restriction enzymes cut double stranded DNA they cleave both strands of DNA and such cuts can be staggered or non-staggered. Experts are tested by Chegg as specialists in their subject area. The resulting DNA fragments are known as restriction fragments.
This process does. 1 2 3 and 4. After digestion of each of these fragments with HindII you find that fragment 3 yields two subfragments 31 and 32 and that fragment 2 yields three 21 22 and 23.
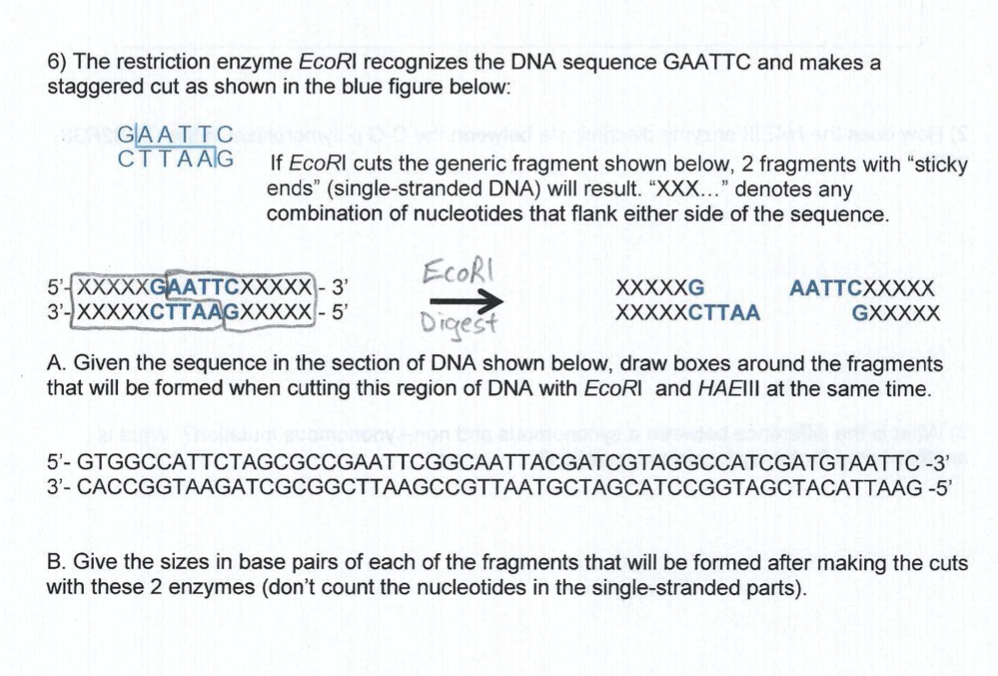
The DNA fragments are separated by electrophoresis a process that involves application of an electric field to cause the DNA fragments to migrate into an agarose gel. We review their content and use your feedback to keep the quality high. After digestion with EcoRI you obtain four fragments.
A restriction enzyme is a DNA-cutting enzyme that recognizes specific sites in DNA. The single strand may generate either at 5 end or to 3 end depending on the enzyme usually the ends have 5 phosphate and 3-hydroxyl ends. Because these overhangs are capable of annealing with complementary overhangs these are called sticky ends Addition of an enzyme called DNA ligase permanently joins the DNA fragments via phosphodiester bonds.
Blunt ends may also be referred to as flush ends. They are naturally produced by bacteria as a defense mechanism against foreign DNA. As a result of this type of cleavage the DNA fragments with blunt ends are generated.
A restriction enzyme or restriction endonuclease is an enzyme that cuts DNA at or near specific recognition nucleotide sequences known as restriction sites. Cohesive ends have unpaired DNA nucleotides on either 5- or 3- strand which are known as overhangs. Because these overhangs are capable of annealing with complementary overhangs we call them sticky ends Adding the enzyme DNA ligase.
Since the restriction sites are symmetrical so that both strands have the same sequence when read in the 5 to 3 direction. Many restriction endonucleases make staggered cuts in the two strands of DNA such that the cut ends have a 2- or 4-base single-stranded overhang. Restriction fragments are the pieces that result after a restriction endonuclease has cut a length of DNA.
Restriction enzymes are also known as restriction endonucleases. However some produce blunt ends. Some enzymes create 5 overhangs and others create 3 overhangs.
Then how many fragments will be generated. The DNA that is cut may have been a small DNA fragment phage DNA a human chromosome or the entire human genome. Restriction enzymes are DNA-cutting enzymes.
Click to see full answer. Why are restriction endonucleases that introduce staggered cuts into DNA especially helpful in generating recombinant DNA molecules. Restriction enzymes are a class of enzymes that cut DNA into fragments based upon recognizing a specific sequence of nucleotides.
However the majority of enzymes make cuts staggered on each strand resulting in a few base pairs of single-stranded DNA at each end of the fragment known as sticky ends. Such ends may be generated by restriction enzymes that break the molecules phosphodiester backbone at specific locations which themselves belong to a larger class of enzymes called exonucleases and endonucleases. Each enzyme has what is known as a recognition.
These overhangs are most often generated by a staggered cut of restriction enzymes.

Restriction Enzymes Create Sticky Or Blunt Dna Ends

Restriction Enzymes Recombinant Dna And Biotechnology Mcat Content

1 12 Restriction Digest With Gel Electrophorisis Biology Libretexts

What Are Staggered Ends In Molecular Biology Quora
Definition Of Cutting And Ligating Dna Molecules Chegg Com

Restriction Enzyme An Overview Sciencedirect Topics

Genetics Ch 19 Molecular Genetic Analysis And Biotechnology Flashcards Quizlet
Solved All Fragments Generated By Staggered Cut By Chegg Com

Solved 6 The Restriction Enzyme Ecori Recognizes The Dna Chegg Com
Restriction Enzymes 101 Nordic Biosite

Solved Restriction Endonucleases Can Cut Dna Into Fragments Chegg Com

Restriction Enzymes Recombinant Dna And Biotechnology Mcat Content

Type Of Dna Ends Generated By Restriction Enzymes Representative Download Scientific Diagram
Restriction Enzymes Endonucleases Methylation Palindromic Sequence

Ppt 1 Restriction Enzyme Kacang Panjang Academia Edu

Restriction Endonucleases An Overview Sciencedirect Topics
Staggered Cut Molecular Biology

Can Autoradiography Distinguish Between Sticky And Blunt Ended Cuts

Type Of Dna Ends Generated By Restriction Enzymes Representative Download Scientific Diagram
Comments
Post a Comment